Diversity of bacterial symbionts in weevils
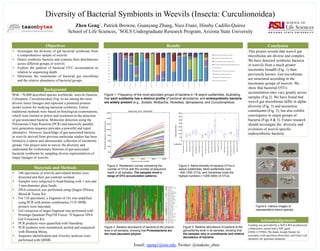
- 1. Zhen Geng^, Patrick Browne, Guanyang Zhang, Nico Franz, Hinsby Cadillo-Quiroz School of Life Sciences, ^SOLS Undergraduate Research Program, Arizona State University Diversity of Bacterial Symbionts in Weevils (Insecta: Curculionoidea) • 246 specimens of weevils and related beetles were dissected and their gut contents isolated. • Samples were subjected to bead-beating with 1 mm and 3 mm-diameter glass beads. • DNA extraction was performed using Qiagen DNeasy Blood & Tissue Kit. • For 110 specimens, a fragment of 16s was amplified using PCR with primer combination 515F-909R; primers were barcoded. • Gel extraction of target fragment was performed with Promega Quantum PrepTM Freeze ‘N Squeeze DNA Gel Extraction Kit. • PCR products were quantified with Nanodrop. • PCR products were normalized, pooled and sequenced with Illumina Miseq. • Sequence identification and diversity analyses were performed with QIIME. Materials and Methods With ~70,000 described species worldwide, weevils (Insecta: Coleoptera: Curculionoidea) (Fig. 6) are among the most diverse insect lineages and represent a potential premier model system for studying bacterial symbiosis. Earlier traditional methods were based on histological examinations, which were limited in power and resolution in the detection of gut-associated bacteria. Molecular detection using the Polymerase Chain Reaction (PCR) and massively parallel, next generation sequence provides a powerful and rapid alternative. However, knowledge of gut-associated bacteria in weevils derived from previous molecular studies has been limited to a sparse and idiosyncratic collection of taxonomic groups. Our project aims to survey the diversity and understand the evolutionary histories of gut-associated bacterial symbionts by sampling diverse representatives of major lineages of weevils. Background • Investigate the diversity of gut bacterial symbionts from a comprehensive sample of weevils. • Detect symbiotic bacteria and examine their distributions across different groups of weevils. • Explore the patterns of bacterial OTU accumulation in relation to sequencing depth. • Determine the constituents of bacterial gut microbiota and the relative abundance of bacterial groups. Objectives Figure 6. Habitus images of representative weevil species Conclusion This project reveals that weevil gut microbiotas are diverse and complex. We have detected symbiotic bacteria in weevils from a much greater taxonomic breadth (Fig. 1) than previously known. Gut microbiotas are structured according to the taxonomic groups of weevils. We show that bacterial OTUs accumulation rates vary greatly across samples (Fig.2). We have found that weevil gut microbiotas differ in alpha- diversity (Fig. 3) and taxonomic constituents (Fig. 5), but also exhibit convergence to major groups of bacteria (Figs 4 & 5). Future research should investigate the diversity and evolution of weevil-specific endosymbiotic bacteria. Subfamily 0% 10% 20% 30% 40% 50% 60% 70% 80% 90% 100% Anthribinae Apioninae Baridinae Ceutorhynchinae Conoderinae Cossoninae Cryptorhynchinae Curculioninae EnDminae Erirhininae Hylobiinae Hyperinae Lixinae MolyDnae Orthognathinae Rhyncophorinae Trachelizinae unplaced Erwinia (Enterobacteriaceae) Sodalis (Enterobacteriaceae) Entomoplasmatales (Mollicutes) Comamonadaceae (Betaproteobacteria) Chloroplast Enterococcus (Enterococcaceae) Wolbachia (RickeSsiaceae) RickeSsia (RickeSsiaceae) Enterobacteriaceae Results Figure 1. Frequency of the most abundant groups of bacteria in 18 weevil subfamilies, illustrating that each subfamily has a distinct profile of bacterial abundance, and endosymbiotic bacteria are widely present (e.g., Sodalis, Wolbachia, Rickettsia, Spiroplasma, and Curculioniphilus). Figure 4. Relative abundance of bacteria at the phylum- level in all samples, showing that Proteobacteria are the most abundant phylum. Figure 2. Rarefaction curves comparing the number of OTUs with the number of sequence reads in all samples. The samples show a range of OTU accumulation patterns. Figure 5. Relative abundance of bacteria at the genus/family-level in all samples, showing that the samples vary in constituents and relative abundance of bacteria. Figure 3. Alpha-diversity of bacteria OTUs in weevil subfamilies. Most subfamilies host ~500-1500 OTUs, and Hylobiinae hosts the highest numbers (~1200-3000) of OTUs. Subfamily Observed OTUs Acknowledgements Funding was provided by a SOLS RTI postdoctoral collaborative award and a NSF grant (DEB-1155984). We thank Joseph Hunter for assistance with specimen dissection, and Franz Lab members for specimen donation. Email: zgeng1@asu.edu, Twitter: @makoto_zhen
