Protocol for maximum parsimony
•Descargar como DOCX, PDF•
0 recomendaciones•438 vistas
it's a protocol for Maximum Parsimony method using MEGA software...
Denunciar
Compartir
Denunciar
Compartir
Recomendados
Associate in Applied Science Network Administration and Information Security

Associate in Applied Science Network Administration and Information SecurityThe Noyes Home For Children
Recomendados
Associate in Applied Science Network Administration and Information Security

Associate in Applied Science Network Administration and Information SecurityThe Noyes Home For Children
Introduction to PaleoWeb by Arwen Vaughan, Rothwell - 2014 PaleoGIS & PaleoCl...

Introduction to PaleoWeb by Arwen Vaughan, Rothwell - 2014 PaleoGIS & PaleoCl...The Rothwell Group, L.P.
USPS® Forced Meter Migration - How to Know if Your Postage Meter Will Soon be...

USPS® Forced Meter Migration - How to Know if Your Postage Meter Will Soon be...Postal Advocate Inc.
Más contenido relacionado
Destacado
Introduction to PaleoWeb by Arwen Vaughan, Rothwell - 2014 PaleoGIS & PaleoCl...

Introduction to PaleoWeb by Arwen Vaughan, Rothwell - 2014 PaleoGIS & PaleoCl...The Rothwell Group, L.P.
Destacado (12)
4 questions linked in an effective teaching for an effective learning

4 questions linked in an effective teaching for an effective learning
Venta B2B Socialmente Responsable, taller Cámara Madrid v 5.04.2016

Venta B2B Socialmente Responsable, taller Cámara Madrid v 5.04.2016
Introduction to PaleoWeb by Arwen Vaughan, Rothwell - 2014 PaleoGIS & PaleoCl...

Introduction to PaleoWeb by Arwen Vaughan, Rothwell - 2014 PaleoGIS & PaleoCl...
Último
USPS® Forced Meter Migration - How to Know if Your Postage Meter Will Soon be...

USPS® Forced Meter Migration - How to Know if Your Postage Meter Will Soon be...Postal Advocate Inc.
Último (20)
INCLUSIVE EDUCATION PRACTICES FOR TEACHERS AND TRAINERS.pptx

INCLUSIVE EDUCATION PRACTICES FOR TEACHERS AND TRAINERS.pptx
Influencing policy (training slides from Fast Track Impact)

Influencing policy (training slides from Fast Track Impact)
ENG 5 Q4 WEEk 1 DAY 1 Restate sentences heard in one’s own words. Use appropr...

ENG 5 Q4 WEEk 1 DAY 1 Restate sentences heard in one’s own words. Use appropr...
USPS® Forced Meter Migration - How to Know if Your Postage Meter Will Soon be...

USPS® Forced Meter Migration - How to Know if Your Postage Meter Will Soon be...
Millenials and Fillennials (Ethical Challenge and Responses).pptx

Millenials and Fillennials (Ethical Challenge and Responses).pptx
AUDIENCE THEORY -CULTIVATION THEORY - GERBNER.pptx

AUDIENCE THEORY -CULTIVATION THEORY - GERBNER.pptx
MULTIDISCIPLINRY NATURE OF THE ENVIRONMENTAL STUDIES.pptx

MULTIDISCIPLINRY NATURE OF THE ENVIRONMENTAL STUDIES.pptx
Student Profile Sample - We help schools to connect the data they have, with ...

Student Profile Sample - We help schools to connect the data they have, with ...
Protocol for maximum parsimony
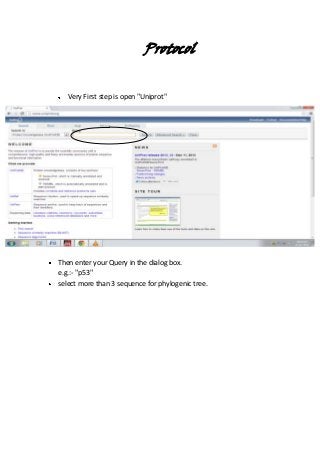
- 1. Protocol Very First step is open "Uniprot" Then enter your Query in the dialog box. e.g.:- "p53" select more than 3 sequence for phylogenic tree.
- 2. here 10 sequence selected. Then Retrieve for download sequence. here take "fasta" format for download
- 3. Then open "MEGA" in this open file > open a file/session open fasta format file where you save.... then "align"
- 4. Select all sequence by ctrl+A than from the menu bar select Alignment > Align by clastalW Again from the menu bar select Data > Export Alignment > MEGA Format
- 5. than close the MEGA & open again. from the menu bar select file > open a File/Session this time open that which you are save as mega format not fasta format. then select phylogeny > Construct / Test Maximum Parsimony Tree(s) So the phylogenic tree saw as follow:-
