Suppdata
•
0 recomendaciones•198 vistas
infromacion suplementaria
Denunciar
Compartir
Denunciar
Compartir
Descargar para leer sin conexión
Recomendados
Recomendados
Más contenido relacionado
La actualidad más candente
La actualidad más candente (13)
Lior schenk - pkar subunits exhibit differential localizations and functions...

Lior schenk - pkar subunits exhibit differential localizations and functions...
Candida antarctica-lipase-b-liquid-calb-recombinant-product-information-sheet

Candida antarctica-lipase-b-liquid-calb-recombinant-product-information-sheet
Similar a Suppdata
Similar a Suppdata (7)
Multipex reverse transcription for detection of viruses

Multipex reverse transcription for detection of viruses
Generation of MRP2 Efflux Transporter Knock-Out in HepaRG Cell Line

Generation of MRP2 Efflux Transporter Knock-Out in HepaRG Cell Line
Último
💉💊+971581248768>> SAFE AND ORIGINAL ABORTION PILLS FOR SALE IN DUBAI AND ABUDHABI}}+971581248768
+971581248768 Mtp-Kit (500MG) Prices » Dubai [(+971581248768**)] Abortion Pills For Sale In Dubai, UAE, Mifepristone and Misoprostol Tablets Available In Dubai, UAE CONTACT DR.Maya Whatsapp +971581248768 We Have Abortion Pills / Cytotec Tablets /Mifegest Kit Available in Dubai, Sharjah, Abudhabi, Ajman, Alain, Fujairah, Ras Al Khaimah, Umm Al Quwain, UAE, Buy cytotec in Dubai +971581248768''''Abortion Pills near me DUBAI | ABU DHABI|UAE. Price of Misoprostol, Cytotec” +971581248768' Dr.DEEM ''BUY ABORTION PILLS MIFEGEST KIT, MISOPROTONE, CYTOTEC PILLS IN DUBAI, ABU DHABI,UAE'' Contact me now via What's App…… abortion Pills Cytotec also available Oman Qatar Doha Saudi Arabia Bahrain Above all, Cytotec Abortion Pills are Available In Dubai / UAE, you will be very happy to do abortion in Dubai we are providing cytotec 200mg abortion pill in Dubai, UAE. Medication abortion offers an alternative to Surgical Abortion for women in the early weeks of pregnancy. We only offer abortion pills from 1 week-6 Months. We then advise you to use surgery if its beyond 6 months. Our Abu Dhabi, Ajman, Al Ain, Dubai, Fujairah, Ras Al Khaimah (RAK), Sharjah, Umm Al Quwain (UAQ) United Arab Emirates Abortion Clinic provides the safest and most advanced techniques for providing non-surgical, medical and surgical abortion methods for early through late second trimester, including the Abortion By Pill Procedure (RU 486, Mifeprex, Mifepristone, early options French Abortion Pill), Tamoxifen, Methotrexate and Cytotec (Misoprostol). The Abu Dhabi, United Arab Emirates Abortion Clinic performs Same Day Abortion Procedure using medications that are taken on the first day of the office visit and will cause the abortion to occur generally within 4 to 6 hours (as early as 30 minutes) for patients who are 3 to 12 weeks pregnant. When Mifepristone and Misoprostol are used, 50% of patients complete in 4 to 6 hours; 75% to 80% in 12 hours; and 90% in 24 hours. We use a regimen that allows for completion without the need for surgery 99% of the time. All advanced second trimester and late term pregnancies at our Tampa clinic (17 to 24 weeks or greater) can be completed within 24 hours or less 99% of the time without the need surgery. The procedure is completed with minimal to no complications. Our Women's Health Center located in Abu Dhabi, United Arab Emirates, uses the latest medications for medical abortions (RU-486, Mifeprex, Mifegyne, Mifepristone, early options French abortion pill), Methotrexate and Cytotec (Misoprostol). The safety standards of our Abu Dhabi, United Arab Emirates Abortion Doctors remain unparalleled. They consistently maintain the lowest complication rates throughout the nation. Our Physicians and staff are always available to answer questions and care for women in one of the most difficult times in their lives. The decision to have an abortion at the Abortion Cl+971581248768>> SAFE AND ORIGINAL ABORTION PILLS FOR SALE IN DUBAI AND ABUDHA...

+971581248768>> SAFE AND ORIGINAL ABORTION PILLS FOR SALE IN DUBAI AND ABUDHA...?#DUbAI#??##{{(☎️+971_581248768%)**%*]'#abortion pills for sale in dubai@
Último (20)
Apidays New York 2024 - Passkeys: Developing APIs to enable passwordless auth...

Apidays New York 2024 - Passkeys: Developing APIs to enable passwordless auth...
Apidays New York 2024 - Accelerating FinTech Innovation by Vasa Krishnan, Fin...

Apidays New York 2024 - Accelerating FinTech Innovation by Vasa Krishnan, Fin...
+971581248768>> SAFE AND ORIGINAL ABORTION PILLS FOR SALE IN DUBAI AND ABUDHA...

+971581248768>> SAFE AND ORIGINAL ABORTION PILLS FOR SALE IN DUBAI AND ABUDHA...
Boost Fertility New Invention Ups Success Rates.pdf

Boost Fertility New Invention Ups Success Rates.pdf
Apidays New York 2024 - The value of a flexible API Management solution for O...

Apidays New York 2024 - The value of a flexible API Management solution for O...
ProductAnonymous-April2024-WinProductDiscovery-MelissaKlemke

ProductAnonymous-April2024-WinProductDiscovery-MelissaKlemke
2024: Domino Containers - The Next Step. News from the Domino Container commu...

2024: Domino Containers - The Next Step. News from the Domino Container commu...
Rising Above_ Dubai Floods and the Fortitude of Dubai International Airport.pdf

Rising Above_ Dubai Floods and the Fortitude of Dubai International Airport.pdf
Apidays New York 2024 - APIs in 2030: The Risk of Technological Sleepwalk by ...

Apidays New York 2024 - APIs in 2030: The Risk of Technological Sleepwalk by ...
Polkadot JAM Slides - Token2049 - By Dr. Gavin Wood

Polkadot JAM Slides - Token2049 - By Dr. Gavin Wood
Emergent Methods: Multi-lingual narrative tracking in the news - real-time ex...

Emergent Methods: Multi-lingual narrative tracking in the news - real-time ex...
How to Troubleshoot Apps for the Modern Connected Worker

How to Troubleshoot Apps for the Modern Connected Worker
Connector Corner: Accelerate revenue generation using UiPath API-centric busi...

Connector Corner: Accelerate revenue generation using UiPath API-centric busi...
DEV meet-up UiPath Document Understanding May 7 2024 Amsterdam

DEV meet-up UiPath Document Understanding May 7 2024 Amsterdam
Suppdata
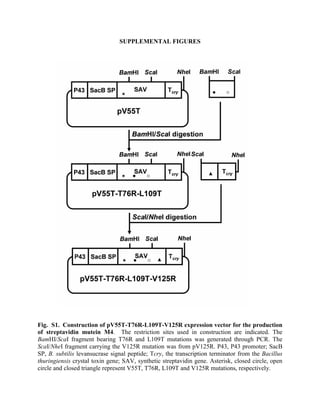
- 1. SUPPLEMENTAL FIGURES Fig. S1. Construction of pV55T-T76R-L109T-V125R expression vector for the production of streptavidin mutein M4. The restriction sites used in construction are indicated. The BamHI/ScaI fragment bearing T76R and L109T mutations was generated through PCR. The ScaI/NheI fragment carrying the V125R mutation was from pV125R. P43, P43 promoter; SacB SP, B. subtilis levansucrase signal peptide; Tcry, the transcription terminator from the Bacillus thuringiensis crystal toxin gene; SAV, synthetic streptavidin gene. Asterisk, closed circle, open circle and closed triangle represent V55T, T76R, L109T and V125R mutations, respectively.
- 2. Fig. S2. Gel filtration analyses of streptavidin muteins. A. Determination of apparent molecular mass of streptavidin muteins in the absence of biotin using Bio-Prep SE 100/17 gel filtration column. Bovine gamma-globulin (158 kDa), bovine serum albumin (66 kDa), ovalbumin (44 kDa), myoglobin (17 kDa) and vitamin B-12 (1.35 kDa) were used as molecular mass markers (closed circles) to calibrate the column. The logarithm of molecular mass was plotted versus Kav (Kav = (Ve-Vo)/(Vt-Vo); Ve, elution volume; Vo, void volume; Vt, bed volume). Positions of wild-type streptavidin (wt) and streptavidin muteins are represented by solid squares. B. Elution profile of M4 mutein (loaded at 2 mg/ml) on the gel filtration column. Fig. S3. Kinetic parameters for the interaction between M4 streptavidin mutein and biotin as monitored by BIAcore biosensor. A. On-rate (ka) determination. ka is derived from association analysis using a plot of ks versus protein concentration (35) B. Off-rate (kd) determination. kd is the slope of the plot of ln(Ro/Rt) versus (t-to) where Ro is the response at an arbitrary starting time to; Rt is the response at time t.
