JBEI Highlights June 2020
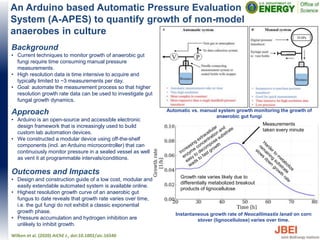
- 1. An Arduino based Automatic Pressure Evaluation System (A-APES) to quantify growth of non-model anaerobes in culture Background • Current techniques to monitor growth of anaerobic gut fungi require time consuming manual pressure measurements. • High resolution data is time intensive to acquire and typically limited to ~3 measurements per day. • Goal: automate the measurement process so that higher resolution growth rate data can be used to investigate gut fungal growth dynamics. Wilken et al. (2020) AIChE J., doi:10.1002/aic.16540 Instantaneous growth rate of Neocallimastix lanati on corn stover (lignocellulose) varies over time. Approach • Arduino is an open-source and accessible electronic design framework that is increasingly used to build custom lab automation devices. • We constructed a modular device using off-the-shelf components (incl. an Arduino microcontroller) that can continuously monitor pressure in a sealed vessel as well as vent it at programmable intervals/conditions. Outcomes and Impacts • Design and construction guide of a low cost, modular and easily extendable automated system is available online. • Highest resolution growth curve of an anaerobic gut fungus to date reveals that growth rate varies over time, i.e. the gut fungi do not exhibit a classic exponential growth phase. • Pressure accumulation and hydrogen inhibition are unlikely to inhibit growth. Automatic vs. manual system growth monitoring the growth of anaerobic gut fungi Measurements taken every minute Growth rate varies likely due to differentially metabolized breakout products of lignocellulose
- 2. Visualizing pectin polymer-polymer entanglement produced by interfacial water movement Background • Pectin polymers have bioadhesive properties that make them interesting for applications e.g. in wound healing, drug delivery and as mesothelial sealants. However, the physico-chemical properties are not fully understood, and a more predictive understanding of the structure-function relationships is required. • In this report, we investigated the physical conditions for creating pectin polymer-polymer (homopolymer) entanglement. Approach • The role of water movement in creating pectin entanglement was investigated by placing water droplets between two glass phase films and compressing the films at variable probe velocities. • Adhesion strength and polymer entanglement were determined as a function of time and compression strength. • This work was a collaboration with Harvard Medical School, Center for Bioenergy Innovation (CBI) and two German universities. Outcomes and Impacts • Slow probe velocity (0.5 mm/sec) demonstrated no evidence of stranding or polymer entanglement. • In contrast, fast probe velocity (5 mm/sec) resulted in 1) an increase in peak adhesion strength, 2) a progressive debonding curve, and 3) increased work of cohesion (p < .001). • Strong adhesion developed rapidly. • We conclude that water movement can supply the motive force for the rapid chain entanglement between pectin films. • The findings are important for developing pectin films and protocols for use as mesothelial sealants. • Pectin is a biomass component and its use in biomedical applications could increase the value of bioenergy crops. Pierce et al. (2020) Carbohydrate Polymers, doi: 10.1016/j.carbpol.2020.116618 Schematic of the adhesion test. The probe moves down at a set velocity until a set force is encountered and is then held at that force. After a set time the probe is moved up at 0.2 mm/sec and adhesion measured. Adhesion develops rapidly with at least 75% of maximum observed after 20 sec. Entanglement is evidenced when adhesion is not instantaneously relaxed but persists as the probe moves away (arrow).
- 3. Do cell wall esters facilitate forest response to climate? Background • Stresses associated with environmental change impact plant growth, development and mortality and ecological succession. • Carbon stocks and biosphere-atmosphere fluxes of CO2, water and VOCs (volatile organic compounds including methanol and acetic acid) are dramatically changing in response to climate factors. • Understanding what drives the forest response to climate change is important for predicting future terrestrial ecosystems responses. • Little is known about the function of cell wall esterification in trees, but pectin methylation is suggested to be important for gas exchange via stomatal control and xylan acetylation is required for proper xylem functioning. • Cell wall esters are also a target for engineering improved biomass for biofuels and renewable chemicals as they can impact downstream fermentation. Dewhirst et al. (2020) Trends in Plant Science, doi: 10.1016/j.tplants.2020.05.011 Integration of cell wall esters with atmospheric emissions and primary carbon metabolism Outcomes and Impacts • In this review, we suggest that cell wall methyl and O-acetyl esters play an important role mediating responses to environmental change, linking modifications in cell wall structure and function with atmospheric fluxes of methanol, acetic acid, CO2 and water. • In addition to being released into the atmosphere, methanol and acetate derived from cell wall esters has the potential to be re- integrated into central carbon metabolism via photorespiration. These cell wall derived compounds could represent an alternative carbon source and play a role in thermal tolerance. • We highlight the need for further interdisciplinary research linking cell wall biochemistry and metabolism with plant physiology and biosphere-atmosphere gas exchange. Climate change related forest dynamics associated with cell wall-derived methanol and acetic acid
- 4. Purification and characterization of a native lytic polysaccharide monooxygenase from Thermoascus aurantiacus Background • Lytic polysaccharide monooxygenases (LPMOs) have emerged as key proteins for depolymerization of cellulose. These copper-containing enzymes oxidize C-1 and/or C-4 bonds in cellulose, promoting increased hydrolysis of the oxidized cellulose chains. • The LPMO from Thermoascus aurantiacus, a thermophilic ascomycete fungus, has been extensively studied and has served as a model LPMO. Fritsche et al. (2020) Biotechnology Letters, doi: 10.1007/s10529-020-02942-w Approach • LPMOs are usually expressed in heterologous host, here a method was developed to purify the LPMO from T. aurantiacus filtrates along with a cellobiohydrolase and endoglucanase. • Also, the effect of the LPMO in T. aurantiacus filtrates was measured by comparing saccharifications with CTec2, which has an LPMO in a commercial cellulase mixture. Outcomes and Impacts • The activity of the purified LPMO was measured with a colorimetric assay that established the Topt of the native LPMO at 60℃. • Quantification of the amounts of cellulases present in the T. aurantiacus culture filtrate established that the LPMO was the most abundant protein in the culture supernatants. • The importance of the LPMO to activity of the mixture compared to CTec2 was demonstrated by saccharifications with Avicel and acid-pretreated corn stover. Purified cellulases from T. aurantiacus Effect of ascorbate supplementation on glucose release Avicel Acid-pretreated corn stover
- 5. An iron (II) dependent oxygenase performs the last missing step of plant lysine catabolism Background • Lysine is an essential amino acid, and due to its low abundance in cereals and legumes it is produced on a scale of one million tons a year to supplement global food supply needs. • Engineered plants with increased lysine could be an integral part of a holistic supply chain for bioenergy crops with multiple end uses. • Despite intensive study, plant lysine catabolism beyond the 2- oxoadipate (2OA) intermediate remains unvalidated. Approach • Recently JBEI described a missing step in the D-lysine catabolism of Pseudomonas putida in which 2OA is converted to D-2- hydroxyglutarate (2HG) via hydroxyglutarate synthase (HglS), a DUF1338 family protein. • We obtained the crystal structure of HglS to 1.1 Å resolution in substrate-free form and in complex with 2OA and use this information to characterize and engineer variants in planta. Outcomes and Impacts • We propose a successive decarboxylation and intramolecular hydroxylation mechanism forming 2HG in a Fe(II)- and O2- dependent manner. • An Arabidopsis thaliana HglS homolog coexpresses with known lysine catabolism enzymes, and mutants show phenotypes consistent with disrupted lysine catabolism. • Results suggest DUF1338-containing enzymes catalyze the same biochemical reaction, exerting the same physiological function across bacteria and eukaryotes. • This knowledge can serve as a guide in engineering plants with increased lysine content. Thompson et al. (2020) Nature Communications, doi:10.1038/s41467-020-16815-3 Ribbon diagram of the HglS crystal structure. The left image shows the active site entrance, containing the metal cofactor (red sphere) and metal-coordinating residues (orange) located in the central β-sheet protein domain. A 180° rotation on the right shows the overall protein structure. Overlay of holo (green) and substrate-bound (cyan) structures of HglS displaying the enzyme active site and the interaction of Val402 and Ser403 with the 2-oxoadipate substrate.
- 6. Investigation of indigoidine synthetase reveals a conserved active-site base residue of nonribosomal peptide synthetase oxidases Pang et al. (2020) J Am Chem Soc., doi:10.1021/jacs.0c04328 Mechanistic insight into NRPS Ox domains. (a) The key active-site residue, Tyr 665, of the BpsA Ox domain (yellow) was determined by structural alignment with McbC (gray), a RiPP dehydrogenase. The generality of the key Tyr residue was confirmed by (b) multiple sequence alignment of known NRPS Ox domains and (c) biochemical assays of EpoB-Ox, the Ox domain in the PKS− NRPS assembly line for epothilone biosynthesis. (d) Proposed E2-like mechanism for NRPS Ox domains. Background • Nonribosomal peptide synthetases (NRPSs) and some polyketide synthases (PKSs) are modular enzymes that synthesize secondary metabolites that can be valuable bioproducts. • The NRPS oxidase (Ox) domain usually oxidizes thiazoline, derived from L-cysteine by the cyclization (Cy) domain, to thiazole on the growing intermediate. • The lack of a mecha nistic understanding of the NRPS Ox domains, especially for the defining biochemical active-site residue, has hindered its engineering. Approach • To establish a suitable system for this study, we chose BpsA, the indigoidine synthetase from Streptomyces lavendulae, which is readily expressed in several hosts, and reconstituted its activity in vitro. • Screened different variants to determine activity, mechanism and specificity of BpsA. Outcomes and Impacts • Results show that the Ox domain utilizes an active-site base residue, tyrosine 665, to deprotonate a protein-bound L-glutaminyl residue. • The revealed mechanism of NRPS Ox domains echoes the unified paradigm for azole biosynthesis in RiPPs and NRPS peptides. • These findings not only resolve the biosynthetic pathway mediated by indigoidine synthetase but enabled mechanistic insight into NRPS Ox domains to be used in future pathway engineering efforts.
- 7. Construction of a novel dual-inducible duet- expression system for gene (over)expression in Pseudomonas putida Background • Pseudomonas putida is a widely used host for metabolic engineering and synthetic biology. However, the use of P. putida could be expanded by designing new plasmid vectors for the expression of heterologous proteins. Gauttam et al. (2020) Plasmid, doi: 10.1016/j.plasmid.2020.102514 Approach • To widen the scope of expression vectors for gene co- expression studies, a previously established dual-inducible expression vector, pRG_Duet2, developed for Corynebacterium glutamicum has been modified for use in P. putida. • This expression vector, named pRGPDuo2, harbors two origins of replication, ColE1 for replication in E. coli and pRO1600 for replication in P. putida. Outcomes and Impacts • Two multiple cloning sites (MCS1 and MCS2) in pRGPDuo2 are individually controlled by inducible promoters Ptac or PtetR/tetA. • Functional validation of pRGPDuo2 was confirmed by the co- expression of genes for the fluorescent proteins superfolder green fluorescent protein (sfGFP), and red fluorescent protein (RFP). • The expression vector pRGPDuo2 is an attractive addition to the existing repertoire of expression plasmids for expression profiling and adds to the tools available for P. putida metabolic engineering. Plasmid map of duet vector RFP and sfGFP activity of duet vectors