7.1 dna & replication
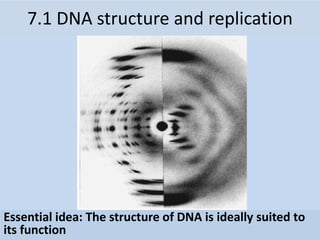
- 1. Essential idea: The structure of DNA is ideally suited to its function 7.1 DNA structure and replication
- 2. Understandings Statement Guidance 7.1 U.1 Nucleosomes help to supercoil the DNA. 7.1 U.2 DNA structure suggested a mechanism for DNA replication 7.1 U.3 DNA polymerases can only add nucleotides to the 3’ end of a primer 7.1 U.4 DNA replication is continuous on the leading strand and discontinuous on the lagging strand. [Details of DNA replication differ between prokaryotes and eukaryotes. Only the prokaryotic system is expected.] 7.1 U.5 DNA replication is carried out by a complex system of enzymes. [The proteins and enzymes involved in DNA replication should include helicase, DNA gyrase, single strand binding proteins, DNA primase and DNA polymerases I and III.] 7.1 U.6 Some regions of DNA do not code for proteins but have other important functions. [The regions of DNA that do not code for proteins should be limited to regulators of gene expression, introns, telomeres and genes for tRNAs.]
- 3. Applications and Skills Statement Utilization 7.1 A.1 Rosalind Franklin’s and Maurice Wilkins’ investigation of DNA structure by X-ray diffraction 7.1 A.2 Use of nucleotides containing deoxyribonucleic acid to stop DNA replication in preparation of samples for base sequencing. 7.1 A.3 Tandem repeats are used in DNA profiling. 7.1 S.1 Analysis of results of the Hershey and Chase experiment providing evidence that DNA is the genetic material. 7.1 S.2 Utilization of molecular visualization software to analyze the association between protein and DNA within a nucleosome
- 4. 7.1 S.1 Analysis of results of the Hershey and Chase experiment providing evidence that DNA is the genetic material. Hershey and Chase Experiments (1952): Definitive proof that DNA rather than Protein carries the hereditary information of life E. Coli bacteriophage: A virus that infects bacteria. Bacteriophages only contain a protein coat (capsid) and DNA. They wanted to find out whether the protein or DNA carried the genetic instructions to make more viruses. They labeled either the viral proteins or DNA: – Protein capsid: Labeled with radioactive sulfur (35S) – DNA: Labeled with radioactive phosphorus (32P) Radioactive labeled viruses were used to infect cells.
- 5. Either Bacteriophage DNA or Proteins Can be Labeled with Radioactive Elements 7.1 S.1 Analysis of results of the Hershey and Chase experiment providing evidence that DNA is the genetic material.
- 6. Hershey Chase Experiment: DNA is Genetic Material 7.1 S.1 Analysis of results of the Hershey and Chase experiment providing evidence that DNA is the genetic material.
- 7. Hershey and Chase Experiments (1952): Bacterial cells that were infected with the two types of bacteriophage, were then spun down into a pellet (centrifuged), and examined. Results: 1. Labeled viral proteins did not enter infected bacteria (found in supernatant). 2. Labeled viral DNA did enter bacteria during viral infection (found in cell pellet). Conclusion: Protein is not necessary to make new viruses. DNA is the molecule that carries the genetic information to make new viruses!!!! 7.1 S.1 Analysis of results of the Hershey and Chase experiment providing evidence that DNA is the genetic material.
- 8. Rosalind Franklin (1950’s) • Worked with Maurice Wilkins • X-ray crystallography = images of DNA • Provided measurements on chemistry of DNA 7.1 A.1 Rosalind Franklin’s and Maurice Wilkins’ investigation of DNA structure by X-ray diffraction James Watson & Francis Crick (1953) • Discovered the double helix by building models to conform to Franklin’s X-ray data and Chargaff’s Rules.
- 9. DNA Double Helix Nitrogenous Base (A,T,G or C) “Rungs of ladder” “Legs of ladder” Phosphate & Sugar Backbone DNA • Two strands coiled called a double helix • Sides made of a pentose sugar Deoxyribose bonded to phosphate (PO4) groups by phosphodiester bonds • Center made of nitrogen bases bonded together by weak hydrogen bonds 7.1 U.2 DNA structure suggested a mechanism for DNA replication
- 10. DNA • Stands for Deoxyribonucleic acid • Made up of subunits called nucleotides • Nucleotide made of: 1. Phosphate group 2. 5-carbon sugar 3. Nitrogenous base 7.1 U.2 DNA structure suggested a mechanism for DNA replication
- 11. DNA Nucleotide O=P-O O Phosphate Group N Nitrogenous base (A, G, C, or T) CH2 O C1 C4 C3 C2 5 Sugar (deoxyribose) O 7.1 U.2 DNA structure suggested a mechanism for DNA replication
- 12. Pentose Sugar • Carbons are numbered clockwise 1’ to 5’ CH2 O C1 C4 C3 C2 5 Sugar (deoxyribose) 7.1 U.2 DNA structure suggested a mechanism for DNA replication
- 14. Antiparallel Strands • One strand of DNA goes from 5’ to 3’ (sugars) • The other strand is opposite in direction going 3’ to 5’ (sugars) 7.1 U.2 DNA structure suggested a mechanism for DNA replication
- 15. Nitrogenous Bases • Double ring PURINES Adenine (A) Guanine (G) • Single ring PYRIMIDINES Thymine (T) Cytosine (C) T or C A or G 7.1 U.2 DNA structure suggested a mechanism for DNA replication
- 16. Base-Pairings • Purines only pair with Pyrimidines • Three hydrogen bonds required to bond Guanine & Cytosine CG 3 H-bonds 7.1 U.2 DNA structure suggested a mechanism for DNA replication
- 17. T A •Two hydrogen bonds are required to bond Adenine & Thymine 7.1 U.2 DNA structure suggested a mechanism for DNA replication
- 18. DNA Replication 7.1 U.3 DNA polymerases can only add nucleotides to the 3’ end of a primer
- 19. Synthesis Phase (S phase) • S phase during interphase of the cell cycle • Nucleus of eukaryotes Mitosis -prophase -metaphase -anaphase -telophase G1 G2 S phase interphase DNA replication takes place in the S phase. Mitosis -prophase -metaphase -anaphase -telophase G1 G2 S phase interphase DNA replication takes place in the S phase. 7.1 U.3 DNA polymerases can only add nucleotides to the 3’ end of a primer
- 20. • DNA replication is very specific to the arrangements of base pairs • In DNA replication, the strands separate – Enzymes use each strand as a template to assemble the new strands DNA REPLICATION Parental molecule of DNA Both parental strands serve as templates Two identical daughter molecules of DNA Nucleotides A 7.1 U.3 DNA polymerases can only add nucleotides to the 3’ end of a primer
- 21. • DNA replication begins at the different origins in the 5’ to 3’ direction at the replication fork. • RNA primase attaches to the DNA and adds a small RNA primer to provide a free 3’ OH starting point since DNA polymerases can only add nucleotides to the 3’ end of a primer • DNA polymerase III adds free nucleotides in the 5’ to 3’ direction in the direction of the replication fork. 7.1 U.3 DNA polymerases can only add nucleotides to the 3’ end of a primer
- 22. • Begins at Origins of Replication • Two strands open forming Replication Forks (Y- shaped region) • New strands grow at the forks Replication Fork Parental DNA Molecule 3’ 5’ 3’ 5’ 7.1 U.3 DNA polymerases can only add nucleotides to the 3’ end of a primer
- 23. DNA Replication • As the 2 DNA strands open at the origin, Replication Bubbles form • Prokaryotes (bacteria) have a single bubble • Eukaryotic chromosomes have MANY bubbles Bubbles Bubbles 7.1 U.4 DNA replication is continuous on the leading strand and discontinuous on the lagging strand. [Details of DNA replication differ between prokaryotes and eukaryotes. Only the prokaryotic system is expected.]
- 24. 7.1 U.4 DNA replication is continuous on the leading strand and discontinuous on the lagging strand. [Details of DNA replication differ between prokaryotes and eukaryotes. Only the prokaryotic system is expected.] DNA replication creates two identical strands with each strand consisting of one new and one old strand (semi-conservative). DNA replication occurs at many different places on the DNA strand called the origins of replication (represented by bubbles along the strand).
- 25. 1. Helicase: unwinds DNA at origins of replication 2. Initiation proteins separate 2 strands forms replication bubble 3. Single-Strand Binding Proteins attach and keep the 2 DNA strands separated and untwisted 4. Primase: puts down RNA primer to start replication 5. DNA polymerase III: adds complimentary bases to leading strand (new DNA is made 5’ 3’) 6. Lagging strand grows in 3’5’ direction by the addition of Okazaki fragments 7. DNA polymerase I: replaces RNA primers with DNA 8. DNA ligase: seals fragments together 7.1 U.5 DNA replication is carried out by a complex system of enzymes. [The proteins and enzymes involved in DNA replication should include helicase, DNA gyrase, single strand binding proteins, DNA primase and DNA polymerases I and III.]
- 26. 7.1 U.5 DNA replication is carried out by a complex system of enzymes. [The proteins and enzymes involved in DNA replication should include helicase, DNA gyrase, single strand binding proteins, DNA primase and DNA polymerases I and III.]
- 27. DNA Replication • DNA polymerase can only add nucleotides to the 3’ end of the DNA • This causes the NEW strand to be built in a 5’ to 3’ direction RNA PrimerDNA Polymerase Nucleotide 5’ 5’ 3’ Direction of Replication 7.1 U.5 DNA replication is carried out by a complex system of enzymes. [The proteins and enzymes involved in DNA replication should include helicase, DNA gyrase, single strand binding proteins, DNA primase and DNA polymerases I and III.]
- 28. Remember HOW the Carbons Are Numbered! O O=P-O O Phosphate Group N Nitrogenous base (A, G, C, or T) CH2 O C1 C4 C3 C2 5 Sugar (deoxyribose) 7.1 U.5 DNA replication is carried out by a complex system of enzymes. [The proteins and enzymes involved in DNA replication should include helicase, DNA gyrase, single strand binding proteins, DNA primase and DNA polymerases I and III.]
- 29. 29 Remember the Strands are Antiparallel P P P O O O 1 2 3 4 5 5 3 3 5 P P P O O O 1 2 3 4 5 5 3 5 3 G C T A
- 30. Synthesis of the New DNA Strands • The Leading Strand is synthesized as a single strand from the point of origin toward the opening replication fork RNA PrimerDNA PolymeraseNucleotides 3’5’ 5’ 7.1 U.5 DNA replication is carried out by a complex system of enzymes. [The proteins and enzymes involved in DNA replication should include helicase, DNA gyrase, single strand binding proteins, DNA primase and DNA polymerases I and III.]
- 31. Synthesis of the New DNA Strands • The Lagging Strand is synthesized discontinuously against overall direction of replication • This strand is made in MANY short segments It is replicated from the replication fork toward the origin RNA Primer Leading Strand DNA Polymerase 5’ 5’ 3’ 3’ Lagging Strand 5’ 5’ 3’ 3’
- 32. Lagging Strand Segments • Okazaki Fragments - series of short segments on the lagging strand • Must be joined together by an enzyme Lagging Strand RNA Primer DNA Polymerase 3’ 3’ 5’ 5’ Okazaki Fragment 7.1 U.5 DNA replication is carried out by a complex system of enzymes. [The proteins and enzymes involved in DNA replication should include helicase, DNA gyrase, single strand binding proteins, DNA primase and DNA polymerases I and III.]
- 33. Joining of Okazaki Fragments • The enzyme Ligase joins the Okazaki fragments together to make one strand Lagging Strand Okazaki Fragment 2 DNA ligase Okazaki Fragment 1 5’ 5’ 3’ 3’ 7.1 U.5 DNA replication is carried out by a complex system of enzymes. [The proteins and enzymes involved in DNA replication should include helicase, DNA gyrase, single strand binding proteins, DNA primase and DNA polymerases I and III.]
- 34. Conservation of base sequence The DNA base sequence is usually read down one side of the molecule or the other. The sequence is usually read with reference to the bases and their corresponding identifying letter. e.g. (1) ATG CTC ATT TTA GGG CCC ATA CTC = 24 bases thus we can write the complementary sequence of the other helix as: (2) TAC GAG TAA AAT CCC GGG TAT GAG In DNA replication (1)will act as the template for a new complementary sequence of bases copy 1: (1)ATG CTC ATT TTA GGG CCC ATA CTC (2)TAC GAG TAA AAT CCC GGG TAT GAG
- 35. 7.1 U.1 Nucleosomes help to supercoil the DNA • A nucleosome consists of DNA wrapped around 8 histone proteins. • The DNA wraps twice around the histone protein core. • Another histone protein is attached to the outside of the DNA strand. It helps maintain the colloidal structure of the nucleosome. • DNA, because of its negative charge is attracted to the positive charge on the amino acids of the histone proteins. • Tails of neighboring histones, link up during chromosomal condensation, causing the nucleosomes to pull closer together. • This is part of the supercoiling process that occurs during mitosis and meiosis • Supercoiling in general helps regulate transcription because only certain areas of the DNA are accessible for the production of mRNA by transcription. This regulates the production of a polypeptide. http://pbil.univlyon1.fr/members/sagot/htdocs/tea m/projects/chromo_net/images/epi.jpg
- 36. 7.1 U.6 Some regions of DNA do not code for proteins but have other important functions. [The regions of DNA that do not code for proteins should be limited to regulators of gene expression, introns, telomeres and genes for tRNAs.] • Regulators such as enhancers are short DNA sequences that regulate protein production or to activate transcription. Regulators such as silencers prevent transcription. The figure below shows how an enhancer might affect gene transcription by bringing distal DNA close to the promoter using coiling or uncoiling with nucleosomes. http://employees.csbsju.edu/hjakubowski/classes/ch331/bind/olbindtransciption.html
- 37. 7.1 U.6 Some regions of DNA do not code for proteins but have other important functions. [The regions of DNA that do not code for proteins should be limited to regulators of gene expression, introns, telomeres and genes for tRNAs.] • There are many areas of DNA containing repetitive sequences, especially in eukaryotic DNA, in humans it makes up between 24-37% of our genome. These repetitive areas usually occurs near the ends of chromosomes. • These non-coded regions are called introns, they must be removed before leaving the nucleus for protein synthesis to take place. • Spliceosome are constructed from RNA and are used in eukaryotic cell to remove these introns. http://www.phschool.com/science/biology_place/bio coach/images/transcription/eusplice.gif
- 38. 7.1 U.6 Some regions of DNA do not code for proteins but have other important functions. [The regions of DNA that do not code for proteins should be limited to regulators of gene expression, introns, telomeres and genes for tRNAs.] • These repetitive sequences called telomeres, protect the DNA during replication. Since enzymes can’t replicate all the way to the end of the chromosome, the parts that aren’t copied are part of the telomeres. This prevents the loss of genes near the end of the chromosomes. Telomeres stained green and red at the ends of the chromosomes http://www.newswise.com/images/uplo ads/2010/09/28/karlsederhr.jpg
- 39. • Dideoxyribonucleotides inhibit DNA polymerase during replication, thereby stopping replication from continuing. • Dideoxyribonucleotides with fluorescent markers, is incorporated into sequences of DNA, to stop replication at the point at which they are added. • This creates different sized fragments with fluorescent markers that can be separated by gel electrophoresis and analyzed by comparing the color of the fluorescence with the fragment length. 7.1 A.2 Use of nucleotides containing deoxyribonucleic acid to stop DNA replication in preparation of samples for base sequencing
- 40. 7.1 A.3 Tandem repeats are used in DNA profiling. • Short tandem repeats (STRs), also known as variable tandem repeats (VNTRs) are regions of noncoding DNA that contain repeats of the same nucleotide sequence. These short repeats show variations between individuals in terms of the number of times the sequences is repeated. • For example, CATACATACATACATACATACATACATA is a STR where the nucleotide sequence CATA is repeated 7 times for one individual. However, in another individual, this tandem repeat could occur only 11 times CATACATACATA CATACATACATACATACATACATACATACATA. These variable tandem repeats are the basis for DNA profiling used in crime scene investigations and genealogical tests (paternity tests). The diagram below shows how the different number of these alleles are used to create a DNA fingerprint of an individual. http://www.intechopen.com/source/html/16506/media/image2.jpg
- 41. 7.1 S.2 Utilization of molecular visualization software to analyze the association between protein and DNA within a nucleosome Use the RCSB Protein Bank to read about nucleosomes and examine Jmol images of them. Article on nucleosomes: http://www.rcsb.org/pdb/101/motm.do?mo mID=7 Jmol visualisation of a nucleosome: http://www.rcsb.org/pdb/explore/jmol.do?s tructureId=1AOI&bionumber=1 Identify the two copies of each histone protein. This can be done by locating the ‘tail of each protein’. The tails of the proteins are involved in regulating gene expression. Suggest how the positive charges help to form the nucleosome with the negatively charged DNA molecule.
